Hello forum,
I want to mask-select some geolocated data across Antarctica with the following basic commands
region=[-180, 180, -90, -60]
spacing=“30m/30m”
df2 = pygmt.select(data=df, resolution=“c”, mask=“s/k”, region=region)
dfbm = pygmt.blockmean(df2, region=region, spacing=spacing)
grd = pygmt.xyz2grd(data=dfbm, region=region, spacing=spacing)
fig.basemap(region=region, projection=“S0/-90/40/20c”, frame=True) # Antarctic
pygmt.makecpt(cmap=“jet”, series=[0, 20], background=“o”)
fig.grdimage(grid=grd)
fig.coast(shorelines=“2p,red”)
fig.plot(x=[135, 165, 165, 135], y=[-76.5, -76.5, -75, -75], pen=“2p,gold”, close=True)
fig.colorbar(position=“JCR+v”, frame=[‘x5’, ‘y+l"Kg m^{2}"’], no_bg=False)
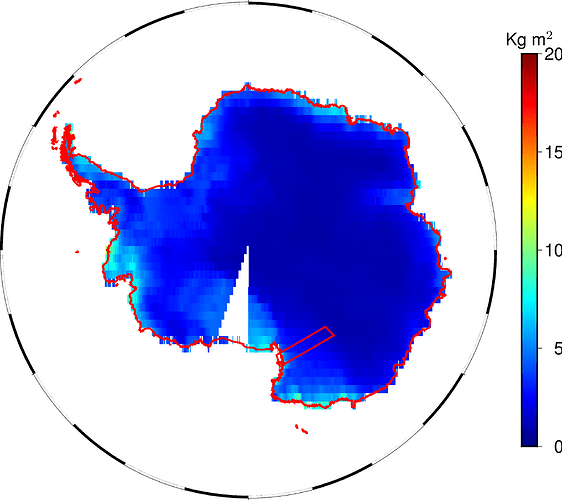
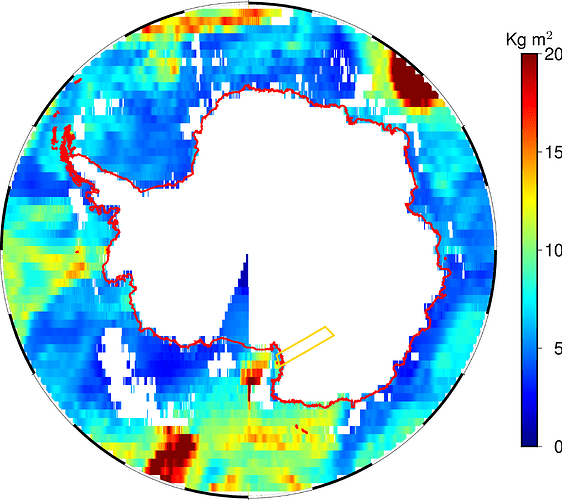
However, some strange pie slice appear for both a land (“s/k”) and an ocean (“k/s”) mask.
Now, that region is the Ross Ice Shelf. It could make sense to exclude it from a land mask if land ice is sorted as “wet” surface, but then the same should hold also for the Ronne Ice Shelf before the Weddel Sea and other minor shelves. And this is not the case.
The shorelines (red coded) indicate that the Ross Ice Shelf is land, indeed.
I am puzzled: is it a select bug, is it a GSHHG problem?
What is your opinion?